DNA recombination and Repair; Protein-DNA Interactions; Nucleic Acid Structure and Dynamics
We employ fluorescence spectroscopic and other biophysical methods to study protein-DNA interactions fundamental to such processes as DNA recombination and repair. One focus of our research program is dynamic mechanisms of recognition in protein-DNA interactions. A particular area of interest is the coding of protein recognition in DNA structure rather than DNA sequence. Our work has also led us to develop FRET mapping methodologies and the use of fluorescent nucleoside analogs such as 6-MI and 6-MAP. In collaboration, we have also used computational methods to complement our spectroscopic studies.
DNA Four-Way Junctions in DNA Repair and Recombination
Many of our studies are focused on DNA four-way or Holliday junctions, a critical intermediate in recombination and repair processes. Our current efforts are focused on the structure and dynamics of this structure with and without proteins bound. 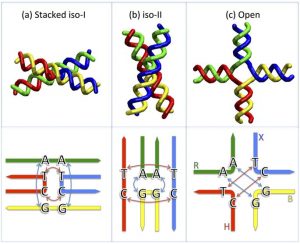 Figure 1 : Idealized schematic of the junction conformations. From: Holliday Junction Thermodynamics and Structure: Coarse-Grained Simulations and Experiments
Figure 1 : Idealized schematic of the junction conformations. From: Holliday Junction Thermodynamics and Structure: Coarse-Grained Simulations and Experiments
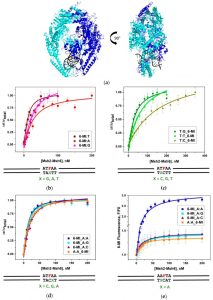
Previously, we have studied the effects of ions and proteins on junction structure. More recently, we have been investigating how the Mut S Homologue (Msh) proteins, Msh2-Msh6 and Msh4-Msh5, interact with the junction structure. Both proteins exhibit high affinity binding to the junction structures. We are currently investigating how the proteins utilize ATP when bound to such structures and protein conformational changes associated with ATP hydrolysis.
Our current work uses these fluorescent probes to examine site-specific base dynamics in the context of protein-DNA interactions. We have successfully applied this methodology to the study of Msh2-Msh6 binding to mismatch DNA and showed that well-recognized mismatches that are efficiently repaired exhibit much higher dynamics in the DNA duplex relative to poorly recognized mismatches and matched DNA.
We have also used this methodology to investigate the amino acid residues involved in the Msh4-Msh5 binding interaction.
Protein-Nucleic Acid Interactions: Architectural Proteins
Nucleotide-binding proteins often play an extremely important role as regulators of genomic function. We are studying the architectural proteins, HU and IHF from E.coli, which bind to the minor groove of DNA through two flexible β-strand regions. These proteins bind to and bend the DNA sculpting it into unique structures requisite for function. We are currently studying the structure and dynamics of the complexes formed between HU and IHF and DNA junctions using x-ray crystallography and single molecule fluorescence methods.
Current lab members: Zane Lombardo, Charya Khun, Nick Taylor, Meera Joshi, Shawn Lin, Daniel Mounier, Shantel Sosa